All published articles of this journal are available on ScienceDirect.
Empirical Analysis of Deep Convolutional Generative Adversarial Network for Ultrasound Image Synthesis
Abstract
Introduction:
Recent research on Generative Adversarial Networks (GANs) in the biomedical field has proven the effectiveness in generating synthetic images of different modalities. Ultrasound imaging is one of the primary imaging modalities for diagnosis in the medical domain. In this paper, we present an empirical analysis of the state-of-the-art Deep Convolutional Generative Adversarial Network (DCGAN) for generating synthetic ultrasound images.
Aims:
This work aims to explore the utilization of deep convolutional generative adversarial networks for the synthesis of ultrasound images and to leverage its capabilities.
Background:
Ultrasound imaging plays a vital role in healthcare for timely diagnosis and treatment. Increasing interest in automated medical image analysis for precise diagnosis has expanded the demand for a large number of ultrasound images. Generative adversarial networks have been proven beneficial for increasing the size of data by generating synthetic images.
Objective:
Our main purpose in generating synthetic ultrasound images is to produce a sufficient amount of ultrasound images with varying representations of a disease.
Methods:
DCGAN has been used to generate synthetic ultrasound images. It is trained on two ultrasound image datasets, namely, the common carotid artery dataset and nerve dataset, which are publicly available on Signal Processing Lab and Kaggle, respectively.
Results:
Results show that good quality synthetic ultrasound images are generated within 100 epochs of training of DCGAN. The quality of synthetic ultrasound images is evaluated using Mean Squared Error (MSE), Peak Signal-to-Noise Ratio (PSNR), and Structural Similarity Index Measure (SSIM). We have also presented some visual representations of the slices of generated images for qualitative comparison.
Conclusion:
Our empirical analysis reveals that synthetic ultrasound image generation using DCGAN is an efficient approach.
Other:
In future work, we plan to compare the quality of images generated through other adversarial methods such as conditional GAN, progressive GAN.
1. INTRODUCTION
Ultrasound imaging is one of the most commonly used primary imaging modalities for prognosis and diagnosis in the medical domain. In comparison to other imaging modalities such as MRI, X-ray, or CT scan, ultrasound is a real-time, radiation-free, robust, and low-cost technique [1, 2]. The increasing demand for computer-aided diagnosis and the innovations in deep neural networks have caused dramatic advances in automated medical image analysis. However, the requirement of a large number of quality images with varying representations of disease hampers the performance of deep neural networks [3, 4]. A deep neural network can learn better only if it uses more number of samples.
Conversely, for rare diseases, it takes many years to collect a vast dataset that is large enough to train the deep model and thereby to prepare a better classifier. Additionally, the low quality of ultrasound images due to speckle noise, low resolution, and contrast mislead the doctors in diagnosis. In such scenarios, image augmentation using GAN has been proven advantageous. It assists not only in producing synthetic images from existing images but also to denoise the input images [4].
Recently, GANs have been considerably used for medical image synthesis. After the invention of GAN, a variety of GANs has been proposed. The random noise vector used as an input to generator in original GAN does not impose any constraint on mutual information between the real image and the synthetic image. To maximize the mutual information, InfoGAN was proposed [5]. It suggests the use of a latent code vector along with a noise vector as an input to the generator network. Conditional GANs have been introduced by conditioning both discriminator and generator for improvement on control over generated data. In a research study [6], Spatially-Conditioned GAN has been used to simulate ultrasound images. An extension of conditional GAN, called Auxiliary Classifier GAN (ACGAN), has been proposed by modifying discriminator architecture to classify samples in addition to the discriminator job. In another study [7], ACGAN has been utilized for the synthesis of liver lesions from CT images. Pix2pix GAN uses conditional GAN with conditional input as an image for image-to-image translation [8]. It uses the discriminator of PatchGAN to produce grid output along with probability for its input. In a research work [9], PatchGAN was used as a discriminator for ultrasound speckle reduction. Pix2pix GAN generates only one synthetic image from one input image, which restricts the diversity of augmented data. To generate a variety of synthetic images, the Pix2Pix model is enhanced by storing network weights as snapshots during training. After each training epoch, the quality of generated images is visually inspected. If images are visually realistic, snapshots are saved to produce different ultrasound images from the single input image. This approach is known as multiple snapshot Pix2Pix approach [10]. For image transformation between two imaging modalities, CycleGAN extracts features of both images and discovers the underlying relationship between them. In a study [11], structured CycleGAN has been used for brain MR-to-CT synthesis. The data distributions of real and synthetic images are matched using Jensen-Shannon (JS) divergence. One of the major drawbacks of JS divergence is vanishing gradients. To avoid vanishing gradient problem, Wasserstein-GAN (WGAN) uses the Earth Mover (ME) distance for measurement of the distance between distributions. While the WGAN model has proven better converge capability, the downside is its slow optimization [12]. Despite wide ultrasound imaging applications, the poor imaging quality and a limited number of available images are major limitations. To overcome these limitations, SpeckleGAN has been designed by adding speckle noise to network feature maps. SpeckleGAN has been used earlier to generate realistic Intra Vascular Ultrasound (IVUS) of the vessel wall [13]. SpeckleGAN improves the overall quality of generated IVUS images compared with a standard GAN model.
In comparison with multi-layer perceptron, CNN has achieved better performance in extracting image features. Due to the fact that CNNs are extremely suitable for image data generation, DCGAN has been proposed as an extension of GAN [14]. Since the invention of DCGAN, researchers have started exploring the capacity of DCGAN to generate a wide variety of medical images. Image augmentation using DCGAN has several applications. Synthetic images can be used as anonymized data to facilitate sharing. This helps to overcome the privacy issues related to the patient’s diagnostic image. GAN-based localization and segmentation of the region of interest (ROI) are helpful for clinical diagnosis of the disease. To assist in the segmentation of lesions in breast ultrasound images, Residual-Dilated-Attention-Gate-UNet (RDAU-NET) has been proposed [16]. This network uses UNet architecture with six residual units and an attention gate to learn lesions by suppressing unnecessary background. The residual units enhance the edge information. To stabilize training, RDAU-NET is combined with WGAN [17]. RDA-UNET-WGAN uses RDAU-NET as the generator, while CNN is used as a discriminator. This hybrid model shows accurate segmentation boundaries as compared to RDAU-net. GAN has been used previously to generate both ultrasound images and their corresponding segmentation images by incorporating dual information in both generator and discriminator [18]. In our approach of DCGAN, we can provide a segmentation mask as a conditional input to generate a segmented image. The discriminator of DCGAN can be utilized to detect abnormalities such as lesions. GAN has also been used for brain lesion detection on MRI [19].
DCGAN has been used to replace simple X-ray and histology images with augmented images for the training of classification models [3]. DCGAN has also been used to improve the performance of liver lesion classification by synthesizing CT images [4]. Besides, it has been revealed that DCGAN performs well in generating brain images by synthesizing MRI [15]. However, to the best of our knowledge, a few attempts have been made in the literature to create artificial ultrasound images using DCGAN. Hence, in our study, we explore and analyze the capability of DCGAN for ultrasound image augmentation.
1.1. Motivation
Though ultrasound image is a crucial diagnostic tool to identify serious and time-sensitive diseases, a dataset with sufficient ultrasound images is not available for several rare diseases to develop an automated diagnostic approach. Therefore, the majority of the existing automated diagnostic applications have acquired images through simulations [4, 7, 24]. However, the images obtained through simulations are not considered appropriate for diagnosis as they involve the life or death decision of humans. Data imbalance is one of the common issues for ultrasound image classification because hospitals generally maintain the records of patients with illness [12]. Moreover, medical records are very sensitive, and it is infeasible and time-consuming to get consent from each patient [3, 7, 9]. Thus, due to all the aforementioned reasons, image augmentation turns out to be an effective alternative to acquire an adequate amount of quality images for the development of an automated approach.
Difficulty in obtaining a large amount of good quality ultrasound images prompted us to explore various image augmentation techniques for generating additional images. The traditional data augmentation methods mainly include translation, rotation, flip, and scaling [21, 22]. However, the application of these methods with random operations eliminates the region of interest from the given image [23]. The invention of GAN in 2014 is a significant breakthrough for artificial data synthesis [24]. GAN estimates the distribution of images to generate a new image that is similar to the original image. The adoption of Convolutional Neural Network (CNN) in computer vision applications is receiving close attention recently. The deep convolutional generative adversarial network is an extension of GAN that uses a deep convolutional network in contrast to the fully-connected network in GAN for both generator and discriminator networks [14]. In this paper, we present an in-depth study and experimental analysis of DCGAN to understand its effectiveness for artificial ultrasound image generation.
The rest of this paper is organised as follows: Section 2 introduces the fundamental concepts of DCGAN. In addition, conventional GAN is discussed in this section. Section 3 describes the details of the datasets and setup of DCGAN used for ultrasound image generation. Results are described and discussed in section 4. Conclusion and future works are specified in section 5.
2. DEEP CONVOLUTIONAL GENERATIVE ADVERSARIAL NETWORK
Conventional GAN is an implicit density estimation generative model that comprises generator and discriminator as two adversarial networks. Fig. (1) gives an overview of the GAN structure.
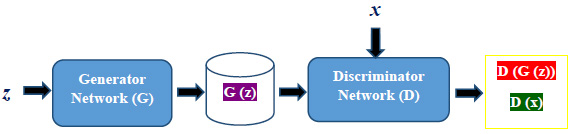
Both generator and discriminator are multi-layer perceptron networks. The generator G with a set of parameters ϴg accepts a noise vector z as an input and produces synthetic images. The discriminator network D with a set of parameters ϴd helps the generator network to optimize the mapping of the distribution of generated images with training images x. D and G play a two-player minimax game [24]. The following equation shows the minimax objective function used to jointly train the generator as well as the discriminator.


ϴg is used to minimize the objective function such that D(G(z)) reaches close to 1. ϴd is used to maximize the objective function such that D(x) reaches close to 1 and D(G(z)) gets close to 0. The output value 0 denotes a fake image, and 1 indicates a real image. Generator and discriminator have been trained alternately in an adversarial manner.
Although GAN performs well, the lack of a heuristic cost function and simultaneous training of both networks result in an unstable situation [7, 8]. Moreover, although it produces good results in generating artificial images, there are still some limitations that hinder GANs’ development. The fundamental limitations include: (1) the mode collapse, (2) non-convergence of model parameters, and (3) diminished gradient. The mode collapse situation occurs when the generator over optimizes, and the discriminator never manages to learn. Due to the mode collapse problem that arises during GAN training, GAN generates synthetic images of limited diversity [15]. The non-convergence of model parameters results in an unstable model that affects the quality of images it produces. When the discriminator successfully rejects synthetic images with high confidence, the generator’s gradient vanishes [24].
To overcome the limitations of GAN, specifically to stabilize training and to avoid mode collapse, deep convolutional GAN incorporates CNN capabilities with the modifications suggested earlier [14]. Batch Normalization and Leaky ReLU activation function are included in DCGAN to improve training stability. DCGAN also replaces all max-pooling layers with strided convolution layers. CNN is used for both generator and discriminator networks to increase the resolution of synthesized images.
The generator uses a random noise vector as an input and feeds it through a fully connected layer and multiple fractionally strided convolution layers to generate fake images. The process of batch normalization and the ReLU activation functions are used at each layer except for the output layer. Tanh activation function is applied to the output layer. The discriminator is a binary classification network that takes an image as an input and produces a scalar probability that represents whether the generated image is real or fake. It consists of multiple convolution layers and a fully connected dense layer. Instead of pooling layers, the convolutions are applied to each convolution layer for reducing the spatial dimensionality. The batch normalization and Leaky ReLU activation function are used at each layer except for the output layer. The sigmoid function is used at the output layer to produce a binary output.
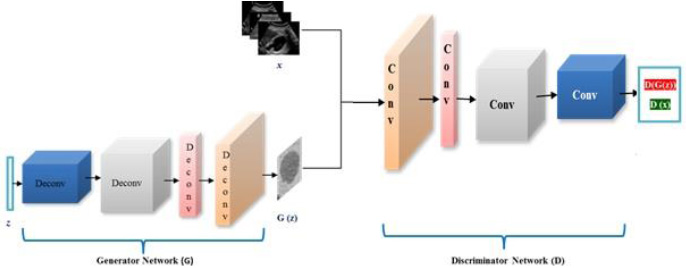
Fig. (2) shows the architecture of DCGAN. The generator network G uses z as an input noise vector and passes it through multiple deconvolutional layers to generate a synthetic ultrasound image represented as G(z). The output of the generator network and training images are fed to discriminator network D to classify them into real and fake images. The discriminator network consists of multiple convolutional layers.
DCGAN generates images in two phases, namely, the training phase and the generation phase. The training phase involves the generator to perform an upsampling operation on a random noise vector for the generation of images similar to real input images. The discriminator helps the generator to improve the quality of synthetic images by classifying the synthetic image as real or fake [13, 14]. Once the training phase finishes, the generator is capable of generating images from randomly selected noise distribution.
3. MATERIALS AND METHODS
3.1. Dataset
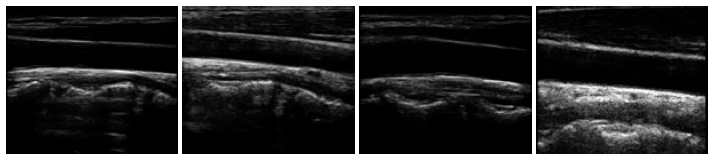
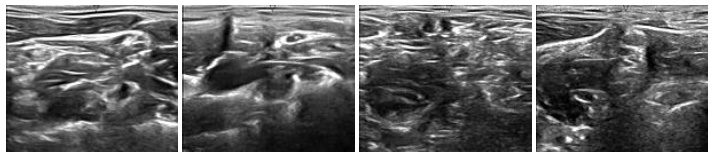
To generate synthetic images, we have experimented with two datasets of ultrasound images that are publicly available on Signal Processing (SP) lab and Kaggle repositories. The dataset on signal processing repository contains a total of 88 B-mode ultrasound images of Common Carotid Artery (CCA), acquired in a longitudinal section. The resolution of these images is 390 x 330 pixels. The nerve image dataset collected from the Kaggle repository contains 5,312 ultrasound images of nerve wherein the brachial plexus is not present. Fig. (3A) shows some sample images from the CCA ultrasound image dataset, and Fig. (3B) shows some sample images from the nerve ultrasound image dataset.
3.2. Experimental Setup
The DCGAN model consists of two convolution neural networks that are trained simultaneously in an adversarial fashion. We conduct experiments with two different CNN architectures with the aim of producing 64 x 64 and 256 x 256 images. In preprocessing, only scaling is applied on the input images to get the range of -1 to 1 for tanh activation. To generate synthetic ultrasound images of size 64 x 64, we follow the architecture of the original DCGAN proposed earlier [14]. To generate synthetic ultrasound images of size 256 x 256, we modify the architecture of the original DCGAN. The following changes are made in the generator network of original DCGAN: 1) For all layers except the output layer, ReLU is replaced by LeakyReLU activation function with a slope of leak = 0.2 to avoid zero gradients and 2) ConvTranspose2D is used for up-sampling to learn parameters of similar ultrasound images. The experimental setting is summarized in Table 1.
Experiments are performed using Pytorch on GPU-supported machines. We use Binary Cross Entropy (BCE) loss function and ADAM optimization with a learning rate of 0.001 for training. Both generator and discriminator networks are trained using stochastic gradient descent (SGD) with a mini-batch size given in Table 1. In each epoch of training, the discriminator followed by the generator has been updated. Loss is adjusted to both the networks separately. For Discriminator update, a batch of real input images is used to calculate the loss value of log(D(x) and a batch of synthetic images generated by the current generator is used to calculate log (1-D(G(z)). The goal of discriminator is to maximize the following function:
Table 1.
| Experiment | Model Architecture | Dataset | Total No. of Images in Dataset | Batch Size |
Output Image Size (in Pixels) |
|---|---|---|---|---|---|
| A | Original DCGAN | CCA dataset from SP Lab | 88 | 8 | 64 x 64 |
| B | Modified DCGAN | CCA dataset from SP Lab | 88 | 8 | 256 x 256 |
| C | Original DCGAN | Nerve dataset from Kaggle | 5312 | 32 | 64 x 64 |
| D | Modified DCGAN | Nerve dataset from Kaggle | 5312 | 32 | 256 x 256 |
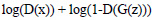
where, x represents real image and z denotes input noise. D and G are discriminator and generator, respectively.
During the training of the generator, the goal is to minimize log (1-D(G(z))) value, which is achieved by maximizing log (D(G(z))).
4. RESULTS AND DISCUSSION
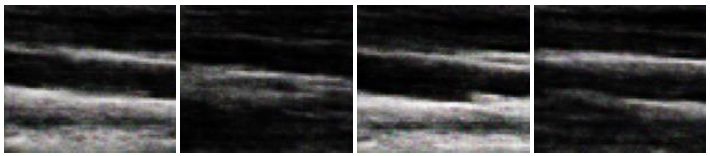
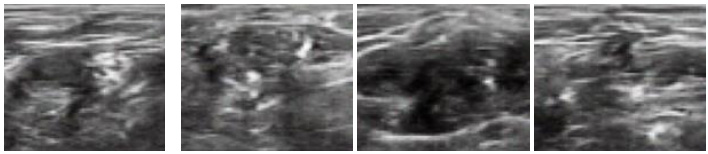
In this section, we report the results obtained using the original DCGAN and modified DCGAN. As discussed in the previous section, original DCGAN is used to generate ultrasound images of size 64 x 64, and modified DCGAN is used to generate ultrasound images of size 256 x 256. Figs. (4 and 5) show the sample synthesized ultrasound images generated using CCA and Nerve dataset. The results illustrate that the DCGAN can produce synthetic ultrasound images of good visual quality. The distribution of grey and white regions in ultrasound images are reproduced in generated images.
The quality of the synthetic ultrasound images is evaluated by calculating Mean Squared Error (MSE), Peak Signal-to-Noise Ratio (PSNR), and Structural Similarity Index Measure (SSIM) [23, 24]. The MSE represents the cumulative squared error between the real and fake images. PSNR measures the similarity between images of a pair wherein the pair consists of a real image and synthetic image. Higher PSNR indicates a higher similarity in the intensity of the synthetic image and real image. SSIM measures the structural similarity of the two images. The higher SSIM represents the higher structural similarity between the synthetic image and the real image [27]. These evaluation measures are defined as follows.
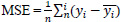
Where n represents the total number of pixels in the image. yi and
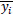 denote real and synthetic images, respectively.
denote real and synthetic images, respectively.
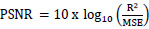
Where R is the maximum fluctuation in the input image. 255 is the default value of R.
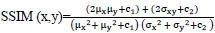
Where, c1 = (k1L)2 and c2 = (k2L)2. x denotes the generated synthetic image and y is the input ultrasound image. μx is the average of x, μy is the average of y, σx2 and σy2 are the variance of x and y, σxy is the covariance between x and y. c1 and c2 prevent the denominator from being zero. L is the dynamic range of the pixel values with k1 = 0.01 and k2 = 0.03.
Table 2 summarizes the values of evaluation metrics for four experiments. The values of evaluation metrics vary marginally in each experiment conducted.
| Experiment | Epochs | MSE mean | MSE median | PSNR mean | PSNR median | SSIM mean | SSIM median |
|---|---|---|---|---|---|---|---|
| A | 100 | 0.19 | 0.13 | 56.25 | 56.90 | 0.98 | 0.99 |
| B | 100 | 0.21 | 0.19 | 55.17 | 55.43 | 0.97 | 0.98 |
| C | 50 | 0.22 | 0.21 | 54.76 | 54.90 | 0.98 | 0.98 |
| D | 50 | 0.26 | 0.23 | 53.97 | 53.11 | 0.96 | 0.97 |
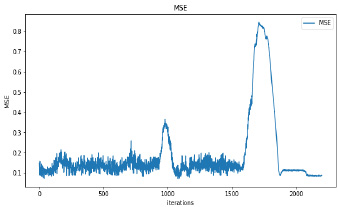
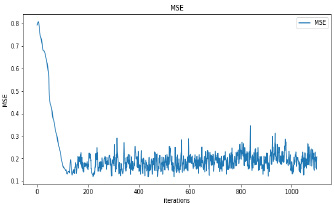
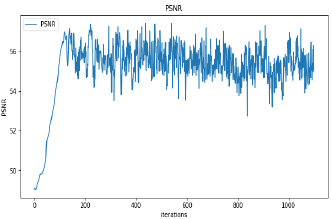
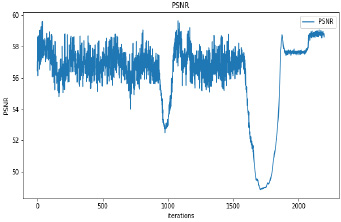
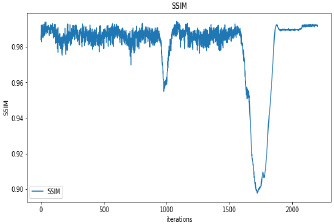
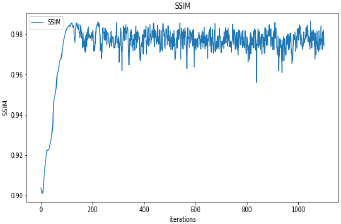
Synthetic images have been generated with 50 and 100 epochs. The experimental results demonstrate that as we increase the number of training epochs, the quality of the synthetic image increases. Fig. (6) presents the plots of MSE, PSNR, and SSIM concerning training iterations for experiments listed in Table 1. For a smaller number of input images, minor underfitting has been observed. For smaller image size, MSE value is increased to its peak value 0.8 at 1500 to 2000 iterations. Interestingly, after 1500 iterations of training, the values of PSNR and SSIM decreased to the lower bound of 50 and 0.90, respectively.
It has been observed that there is no significant difference in PSNR and SSIM values of images synthesized with 50 epochs and 100 epochs. The DCGAN architecture with Leaky ReLU activation for both generator and discriminator stabilizes the convergence within 50 epochs. The synthetic images produced by experiments do not correspond to a real patient; hence, each image’s characteristics are unique to the dataset and thus, possibly increase its variance.
CONCLUSION
The performance of CNN-based ultrasound image analysis relies on both the number of images and the quality of images used for training. GANs have gained tremendous popularity in generating high-quality synthetic images to increase the size of the dataset. Issues such as prerequisite for patient agreement when publishing images and the efforts required to gather images are reduced and resolved by generating realistic synthetic ultrasound images. In this paper, the DCGAN approach has been used for the augmentation of ultrasound images. Our experimental results indicate that high-quality ultrasound images that are indistinguishable from the original images are generated within 100 epochs of training with a maximum average PSNR value of 56.90. The maximum average SSIM value of 0.99 attains structural similarity between generated synthetic images and real images.
In future work, we aim to explore the possibility of performance gain in multimodal ultrasound image generation to represent different pathological cases. We also plan to investigate the combination of other adversarial methods for training instead of dedicated training models.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIAL
Not applicable.
FUNDING
None.
CONFLICTS OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none.


