All published articles of this journal are available on ScienceDirect.
Essentials of Predicting Epileptic Seizures Based on EEG Using Machine Learning: A Review
Abstract
Objective:
Epilepsy is one of the chronic diseases, which requires exceptional attention. The unpredictability of the seizures makes it worse for a person suffering from epilepsy.
Methods:
The challenge to predict seizures using modern machine learning algorithms and computing resources would be a boon to a person with epilepsy and its caregivers. Researchers have shown great interest in the task of epileptic seizure prediction for a few decades. However, the results obtained have not clinical applicability because of the high false-positive ratio. The lack of standard practices in the field of epileptic seizure prediction makes it challenging for novice ones to follow the research. The chances of reproducibility of the result are negligible due to the unavailability of implementation environment-related details, use of standard datasets, and evaluation parameters.
Results:
Work here presents the essential components required for the prediction of epileptic seizures, which includes the basics of epilepsy, its treatment, and the need for seizure prediction algorithms. It also gives a detailed comparative analysis of datasets used by different researchers, tools and technologies used, different machine learning algorithm considerations, and evaluation parameters.
Conclusion:
The main goal of this paper is to synthesize different methodologies for creating a broad view of the state-of-the-art in the field of seizure prediction.
1. INTRODUCTION
Epilepsy is a chronic non-communicable disease of the brain that affects people of all ages. According to the World Health Organization (WHO), around 50 million people suffer from epilepsy worldwide. It is estimated that there are more than 10 million people with epilepsy in India. Globally, an estimated five million people are diagnosed with epilepsy each year. Epilepsy leads to behavioral, health, and economic consequences. Efficient diagnosis and treatment are very much essential for the quality of life of the person with epilepsy and its caregivers.
With the increasing computing availability and storage, machine learning algorithms have been widely explored in the area of epileptic seizure prediction. However, because of the diversified research approaches, dataset considerations, evaluation parameters, and implementation approaches, the reproducibility of the results is limited. The work presented here focuses on the essential aspects of epileptic seizure prediction. The future research approaches may consider the presented comparative analysis of datasets, implementation tools, and methodologies for evolved epileptic seizure prediction methodology.
The organization of this paper is as follows: Section-2 describes the fundamentals of epilepsy and treatment. Section-3 and 4 describe the use of electroencephalography (EEG) for epileptic seizure prediction and the need for algorithms. Section-5 gives the comparative analysis of various approaches for epileptic seizure detection and prediction, followed by Section-6 with the research gaps. Section-7 gives a detailed understanding of datasets used in prior research of epileptic seizure prediction. Section-8 gives a comparative analysis of tools and libraries used for the implementation of epileptic seizure prediction systems, which is followed by the conclusion.
2. EPILEPSY AND TREATMENT
Seizures are due to excessively synchronous and/or excessively intense activity of neuronal circuits in the brain, particularly in the cerebral cortex. In epilepsy, seizures occur spontaneously, repeatedly, and usually suddenly. The unpredictability of seizures represents one of the main disabling features of epilepsy [1]. To date, the causes of epilepsy have not been identified. However, conditions like severe head injury, stroke and blood vessel diseases, tumors, changes in brain structure, and brain infections could provoke seizures [2]. Epileptic seizures can be classified into three groups [3]. Generalized Onset Seizures: These seizures affect both sides of the brain or groups of cells on both sides of the brain at the same time. This term still includes seizure types like tonic-clonic, absence, or atonic. Focal Onset Seizures: It can start in one area or group of cells on one side of the brain. Unknown Onset Seizures: When the beginning of a seizure is not known, it is called an unknown onset seizure.
Epileptic seizures have four different states: the preictal state, which is a state that appears before the seizure begins, the ictal state that begins with the onset of the seizure and ends with an attack; the postictal state that starts after ictal state, and interictal state that starts after the postictal state of first seizure and ends before the start of the preictal state of consecutive seizure [4]. The effect of epilepsy is different in each individual. So, recognizing and diagnosing the type of seizure or epilepsy affecting a person can sometimes be challenging. However, there are a few common ways of testing and determining an epilepsy diagnosis [3]. Various methods, like electroencephalography (EEG), Computerized Tomography (CT) scans, Magnetic Resonance Imaging (MRI), and functional imaging studies, are used to evaluate a person with epilepsy [5, 6]. EEG is the most common method amongst all for the diagnosis and treatment of epilepsy. Persons with epilepsy are generally treated with Anti-Epileptic Drugs (AEDs). A high dosage of AEDs optimally controls epileptic seizures. However, regular consumption of AEDs generally shows side effects like tiredness, headache, dizziness, or blurred vision. It may also lead to behavioral changes in the person with epilepsy [7]. About 20-40% of persons with epilepsy are drug-resistant, i.e., they do not respond to AEDs even though a variety of drugs are available for decades [8, 9], of whom only a small minority can be helped by epilepsy surgery [10].
3. ELECTROENCEPHALOGRAPHY(EEG)
An electroencephalogram (EEG) is the flow of neuronal ionic currents recorded using a pair of electrodes either inside or outside the scalp [11]. The applications of EEG signal processing are brain-computer interface, seizure detection, seizure prediction, schizophrenia detection and classification, diagnosis of Parkinson’s disease, etc. EEG is significantly used in the diagnosis, classification, and treatment of epileptic seizures [8]. If an invasive technique is used to record the EEG signal from inside the skull, it is called intracranial EEG (iEEG). In a non-invasive technique, EEG signals are recorded from the scalp, called scalp EEG (sEEG). EEG waveforms are generally classified into normal and abnormal signals using frequency parameters [12]. EEG signals can be categorised into the following based on frequency: Delta (0.1 - 4 Hz), Theta (4 - 8 Hz), Alpha (8- 13 Hz), Beta (13 - 30 Hz), and Gamma (30 - 100 Hz). Different EEG frequency corresponds to different behavior and mental state of the brain [12]. Due to the high complexity of EEG signals, a single prediction feature can only quantify some of its properties [7].
The popularity of EEG is because of the following advantages: scalp EEG (sEEG) is a non-invasive technique which records the waveforms without much effort or active response by the subject/patient, EEG recording kit is portable and financially affordable, EEG recording devices do not make any noise and no special environment is needed to set it up [11]. Though EEG techniques are widely evolved, there are still a few limitations that need to be addressed. Such as, EEG is prone to low spatial resolution and low signal-to-noise ratio (SNR) [11]. Preprocessing of EEG is also very challenging. To have an artefact-free EEG to extract the control signals, the EEGs have to be restored from the artefacts, such as eye-blinking, electrocardiograms (ECGs), and any other internal or external disturbing effects [13].
4. THE NEED FOR EPILEPTIC SEIZURE PREDICTION ALGORITHMS
With the evolving EEG technology and resource advances, there has been huge interest in the EEG waveform-based research for brain-computer interface (BCI), disease detection, and treatment. Characterization of EEG waveforms plays a vital role in the field of epileptic seizure detection and classification. With the use of sEEG or iEEG signals, certain patterns can be found to detect the preictal state of seizure. The detection of a preictal state would trigger an alarm for the patient or patient’s caregivers to take precautions or medicine beforehand to avoid the ill effect of seizure [7]. Seizure detection methods can be used for the offline analysis of EEG waveforms or seizure-abortion devices. Whereas the seizure prediction system identifies the occurrence of seizure before a certain period called the occurrence period [7].
With rapidly increasing computing power and storage, the availability of EEG data has become easier. Researchers have started using the power of modern machine learning algorithms to improve the results of seizure prediction algorithms [14-27]. Though ample research has already been done, there is no clinical applicability yet [28, 29]. This is because of the sensitivity of the seizure prediction algorithms. One of the challenges in the study of epileptic seizure detection and prediction is to achieve the results to apply the same for clinical applicability [7]. The statistical justification is also desirable for a complete understanding of existing approaches that achieve significant results [7]. For clinical applicability of seizure prediction approaches, the alarm shall trigger prior to a considerable time period. So, the seizure prediction horizon needs to be one of the important evaluation parameters of such systems [7]. Researchers have incorporated various approaches for epileptic seizure prediction. The following sections review the datasets incorporated, tools and libraries used and different algorithms considered for the same. This would help novice ones to explore and improve machine learning-based epileptic seizure prediction algorithms.
5. MACHINE LEARNING ALGORITHMS AND EVALUATION PARAMETERS
The work for epilepsy seizure prediction has evolved drastically since its inception. Based on various methodologies followed and algorithms used, epilepsy seizure prediction can be categorized into four approaches as follows. Also, Fig. (1) shows all four approaches:
- The traditional machine learning approach
- Deep learning approach
- Signal processing approach
- Hybrid approach
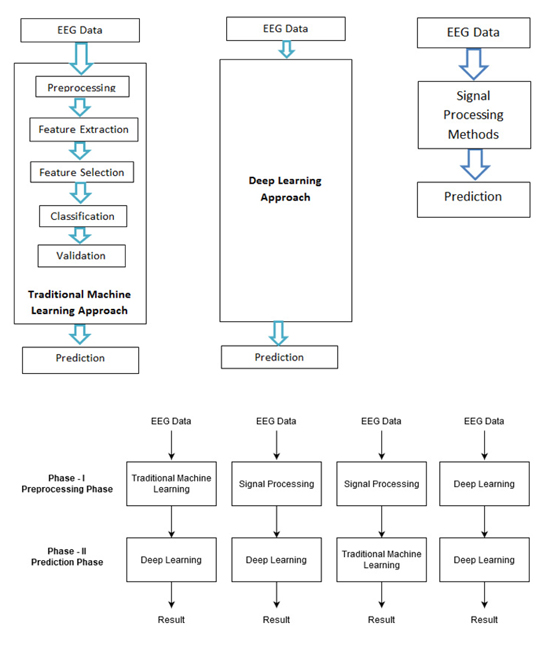
5.1. The Traditional Machine Learning Approach
The traditional machine learning approach includes various stages like preprocessing, feature extraction, feature selection, classification, and validation. Handcrafted features are being used for the detection of the preictal state. A brief description of all the stages is as follows: Data Cleaning, EEG data is nonstationary and prone to artefacts, it requires high preprocessing. This is the phase that takes maximum effort to read, interpret, and clean the noise. Feature Extraction, using different signal processing techniques, features are extracted in time domain, frequency domain, or time and frequency domain. Feature Selection, a huge amount of features is being generated from the feature extraction phase. It is important to find the feature of importance for a specific task. Some of the features considered by researchers are entropy, approximate entropy, Hjorth parameters, spectral moments, mobility, energy, entropy, correlation coefficients, Fast Fourier Transform (FFT), variance, skewness, kurtosis, mean, fractal dimension, frequency band power, peak amplitude, zero crossing, average spectral power, line length, and maximal and minimal values. Classification, the problem of epileptic seizure prediction and detection is actually to differentiate the pre-ictal state of EEG signal from the interictal and ictal state. Different classification algorithms like Support Vector Machine, k-nearest neighbor, Gaussian Naive Bayes, random forest, multi-layer perceptron are used for this purpose, Validation, the hyperparameters are not trained by classification models, it would be predefined. Model performance depends highly on the selection of hyperparameters. The best model amongst various values of hyperparameters is chosen in the validation stage. Prediction is the last phase that performs the task of prediction based on the optimal performing model. Table 1 gives the detailed comparative analysis of various studies conducted using the traditional machine learning approach for epileptic seizure state detection and prediction.
5.2. Deep Learning Approch
Deep learning-based models are end-to-end models, i.e. once EEG signals are given as input, an automated approach for feature extraction and selection is done. Based on the learning and validation, the model would converge and give the prediction results. Another noticeable approach for epileptic seizure prediction is based on signal processing methods. The large amount of data recorded from even a single EEG electrode pair presents a difficult interpretation challenge. Convolutional Neural Network (CNN), Recurrent Neural Network (RNN), and variations of these algorithms have been used by researchers in the past for the task of seizure prediction. This approach is trending because of the availability of resources. Table 2 gives the detailed comparative analysis of various studies conducted using the deep learning approach for epileptic seizure state detection and prediction.
5.3. Signal Processing Approach
In this approach, traditional signal processing methods are used to highly process the data. Signal processing methods are needed to automate signal analysis and interpret the signal phenomena [30]. The prediction stage of this approach uses a basic classification algorithm for signal state classification. Table 3 gives the detailed comparative analysis of various studies conducted using the signal processing approach for epileptic seizure state detection and prediction.
5.4. Hybrid Approach
The combination of all three methods, i.e., traditional machine learning, deep learning, and signal processing leads to a hybrid approach, which takes best of all. Literature provides the data related to the combination of traditional machine learning and deep learning models, deep learning, and signal processing models, machine learning and signal processing methods deep learning methods. References for some hybrid approaches provides the detailed comparative analysis of various studies conducted using the hybrid approach to epileptic seizure state detection and prediction.
To evaluate the performance of various models, different evaluation parameters like sensitivity, specificity, false-positive rate, accuracy, prediction time, AUC, ROC, and F1 score have been considered in the past by other researchers. However, there is no clear consensus regarding which parameter signifies the performance of epileptic seizure detection or prediction algorithms best. Yannick Roy et al. [31] and Alexander Craik et al. [32] have presented a vast comparison of all work done in the field of epileptic seizure prediction. Along with the comparison, various aspects for future research are also mentioned.
6. RESEARCH GAPS
To increase the reliability of seizure prediction results and clinical applicability of the same, evolved models of EEG signal analysis are the need of the time [11]. This section describes some of the improvement areas which could aid in the performance of seizure detection and prediction models, which may lead to clinical applicability. No methods have exhibited both high sensitivity and zero false alarms per hour to achieve the reliability of results [28]. Also, existing machine learning algorithms unnecessarily reduce the number of parameters in feature selection for simplistic classification [28]. The advanced techniques can be used for preprocessing of EEG data to get increased sensitivity of the results [4]. Identification of proper matrices for the evaluation of seizure prediction model is also a big challenge for highly imbalanced data. Most of the work done in the field of epileptic seizure detection and prediction focuses on patient-specific approaches. The concept of domain adaptation and transfer learning can be used for cross-patient research, i.e. generalized models [28]. A better generalization performance between subjects will be necessary to truly make BCIs useful [31]. All prediction methods have been developed and tested on different EEG data pools, making it difficult to compare their performance [101, 119].
| Year | Research Group | Dataset Used | Preprocessing | Feature Extraction | Feature Selection | Classifier Used | Performance |
|---|---|---|---|---|---|---|---|
| 2009 | Theoden Netoff et al. [33] | European Epilepsy Database | Removing artefacts, filtering | Power of spectral bands | N/M | Cost-effective SVM | Sensitivity = 77.8% False positive rate per hour = 0 |
| 2013 | Ning Wang et al. [34] | European Epilepsy Database | N/M | Averaged Instantaneous Envelope (AIE), Averaged Instantaneous Frequency (AIF) | RFE-SVM (Recursive feature elimination - SVM) | SVM | Average Sensitivity = 98.8% Average False alarm per hour = 0.054 AUC = 0.784 |
| 2014 | Peyvand Ghaderyana et al. [35] | European Epilepsy Database | Removing artefacts, normalized power | Spectral bands, statistical moments, median, and Power Spectral Density (PSD), Feature Change Ratio (FCR) | PCA, KNN-based undersampling | SVM | Sensitivity = 100% Average false alarm rate per hour = 0.13 G-mean = 0.97 F-measure = 0.90 |
| 2015 | P. Fergus et al. [36] | CHB-MIT | Delta, theta, alpha, and beta signal bands filtering | Peak Frequency, Median Frequency, variance, root mean squares, sample entropy, skewness, and kurtosis | Linear discriminant analysis backward search | LDC, QDC, UDC, POLYC, LOGLC, KNCC, TREEC, PARZENC, SVC | Best results with KNNC: Sensitivity = 84% Specificity = 85% AUC = 91% Global error = 15% |
| 2015 | Cristian Donos et al. [37] | European Epilepsy Database | N/M | mean, mean absolute deviation, variance, skewness, kurtosis, autocorrelation, line length, power, power ratio | N/M | Random forest classifier | Mean sensitivity = 93.84% Mean detection delays = 3.03s False detections per hour = 0.33/h mean |
| 2015 | Zisheng Zhang et al. [38] | Kaggle - American Epilepsy Society Seizure Prediction Challenge | N/M | PSD features: Relative Spectral Powers, Spectral Power Ratios | CART | SVM | Sensitivity = 100% Mean False Positive (FP) rate = 0.073 FP/hour Mean prediction horizon = 58 minutes AUC = 0.979 |
| 2016 | Bruno Direito et al. [39] | European Epilepsy Database | Butterworth Infinite Impulse Response (IIR) filter to remove the noise |
AR Modeling predictive error, Decorrelation time, Energy Hjorth, Spectral power, Spectral edge, Energy wavelet coefficients, Mean, Variance Skewness, Kurtosis |
N/M | SVM | false predictions per hour = 0.20 sensitivity = 38.47% |
| 2016 | Lung-Chang Lin et al. [40] | Real-time personal dataset | EEG epoch acquisition | Autoregressive modeling predictive error, Decorrelation time, Energy, Entropy, Hjorth, Relative power, Spectral edge, Statistics, Energy of the wavelet coefficients | Correlation based Feature Selection (CFS) approach | SVM | Correctness = 97.50%, Sensitivity = 96.92%, Specificity = 97.78%, Precision = 95.45% |
| 2016 | Khurram I. Qazi et al. [41] | Real-time personal dataset | Seizure, pre-seizure and seizure-free labelling | Energy (E), range (R), standard deviation (SD), the sum of absolute values (SAV), mean absolute values (MAV) and variance (Var) | N/M | SVM, ANN with supervised learning, K means clustering combined with unsupervised learning algorithms | Accuracy = 85 - 90% |
| 2017 | Syed Muhammad Usman et al. [4] | CHB-MIT | Surrogate channel creation, Empirical Mode Decomposition | Statistical features of time-domain and spectral features of frequency domain | N/M | Support Vector Machine, Naïve Bayes, K nearest neighbor | Best results with SVM: Sensitivity = 92.23% and average prediction time = 23.61 minutes |
| 2017 | Han-Tai Shiao [42] | Mayo Clinic dataset | N/M | Spectral features | N/M | SVM | Sensitivity = 90-100% Average False positive rate per day = 0-0.3 |
| 2018 | Yanli Yang et al. [43] | European Epilepsy Database | Filtering and sampling | Permutation entropy | N/M | SVM | Average sensitivity = 94% False prediction rates per hour(FPRh) = 0.111 Average prediction horizon = 61.93min |
| 2019 | Amirhossein Ahmadi et al. [44] | CHB-MIT | N/M | Shannon entropy | Statistical one-sample t-test | SVM,KNN | Sensitivity = 83.8% (SVM & KNN) Specificity = 71% (SVM), 67.8 (KNN) |
| 2019 | Xiashuang Wang et al. [45] | Bonn University Dataset | Digital filtering, removing artefacts, re-referencing, and baseline corrections | Fourier transform, multitaper spectral analysis, PACF, and STFT | N/M | Random forest algorithm based on grid search optimization | Accuracy = 96.7% AUC = 99.0% |
| 2019 | Yuxing Wang et al. [46] | CHB-MIT | Pre-ictal state partition | Wavelet Packet Decomposition (WPD) | N/M | Random forest | Accuracy = 84.8% |
| Year | Research Group | Dataset Used | Preprocessing | Feature Extraction | Feature Selection | Classifier Used | Performance |
|---|---|---|---|---|---|---|---|
| 2016 | Mohammad-Parsa Hosseini et al. [17] | The University of Pennsylvania and the Mayo Clinic | Dimensionality reduction | N/M | N/M | Stacked auto encoder | Accuracy = 0.94 Precision = 0.95 Sensitivity = 0.93 False Positive Rate = 0.05 False Negative Rate = 0.06 |
| 2017 | Sachin Talathi [16] | Bonn University database | N/M | N/M | N/M | Gated Recurrent Unit (GRU) RNNs for seizure detection | Accuracy = 100% Sensitivity = 98% |
| 2017 | U. Rajendra Acharya et al. [47] | Bonn University database | Normalization | N/M | N/M | CNN | Accuracy = 88.67% Sensitivity = 95% Specificity = 90% |
| 2017 | Haidar Khan et al. [48] | 1) Mount Sinai Epilepsy Center dataset 2) CHB-MIT |
N/M | N/M | N/M | Convolutional filters on the wavelet transformation | Sensitivity = 87.8% False prediction rate = 0.142/h |
| 2018 | Isabell Kiral-Kornek et al. [20] | Cook et al.,2013 | Spectrogram generation | N/M | N/M | Deep neural network | Mean sensitivity = 69% Mean time in warning = 27% |
| 2018 | Matthias Eberlein et al. [19] | Kaggle | N/M | N/M | N/M | CNN | AUC = 0.73 |
| 2018 | David Ahmedt-Aristizabal et al. [21] | Bonn University database | N/M | N/M | N/M | LSTM | Accuracy = 95.54% Sensitivity = 91.83 Specificity = 90.50 Precision = 91.50 AUC = 0.9582 |
| 2018 | Nhan Duy Truonga et al. [49] | CHB-MIT, European Epilepsy Database, Kaggle | Short-time Fourier transform, Windowing | N/M | N/M | CNN | Sensitivity = 81.4% - 81.2% - 75% False prediction rate = 0.06/h, 0.16/h, 0.21/h |
| 2018 | Mengni Zhou et al. [50] | European Epilepsy Database, CHB-MIT | Spectrogram generation | N/M | N/M | CNN | Accuracy = 93-97.5% |
| 2018 | R. Schirrmeister et al. [15] | The Temple University Hospital (TUH) EEG Abnormal Corpus |
Artefact removal and resampling | N/M | N/M | CNN | Accuracy = 84.5-85.4% Sensitivity = 75.1-77.3% Specificity = 90.5-94.1% Accuracy = 81.7-82.5% |
| 2019 | Xinghua Yao et al. [51] | CHB-MIT | Bidirectional Long Short-Term Memory (BiLSTM) | N/M | N/M | Softmax function | Sensitivity = 87% Specificity = 88.60% Precision = 88.63% |
| 2019 | Xinghua Yao et al. [23] | CHB-MIT | Independently recurrent neural network (IndRNN) | N/M | N/M | N/M | Sensitivity = 87.3% Specificity = 86.7% Precision = 87.08% F1 score = 87.07% |
| 2019 | Ali Emamia et al. [27] | NTT Medical Center Tokyo | 2-D image construction | N/M | N/M | CNN | Median of detected seizure rate by minutes = 100% False alarm 0.2 per hour |
| 2019 | Ibrahim Aliyu et al. [52] | Bonn University database | Discrete wavelet transform |
RNN-LSTM | Accuracy = 99% | ||
| 2019 | Chien-Liang Liu et al. [53] | Kaggle, CHB-MIT | PCA, FFT, and data augmentation | N/M | N/M | Multi-view CNN | AUC = 0.84 (Kaggle), 0.82-0.89 (CHB-MIT) |
| 2020 | Fabio Pisano et al. [54] | European Epilepsy Database | Manual channel selection, EEG segmentation and data augmentation | N/M | N/M | CNN | Accuracy = 96.39% Specificity = 96.81% Sensitivity = 93.20% Gmean = 89.92-98.83% |
| Year | Research Group | Dataset Used | Preprocessing | Feature Extraction | Feature Selection | Classifier Used | Performance |
|---|---|---|---|---|---|---|---|
| 2012 | James R. Williamson et al. [55] | European Epilepsy Database | Filtering and normalization | High-dimensional feature vectors are extracted from space–delay covariance and correlation matrices | N/M | SVM | Sensitivity = 95-86% AUC = 0.973 |
| 2012 | Mojtaba Bandarabadi et al. [56] | European Epilepsy Database | Filtering and windowing | Normalized spectral power features, relative features using bi-variate approach | normalized difference of the percentiles | SVM | Sensitivity = 76.09% False positive rate per hour = 0.15 No. of sleeted features = 8.75 Seizure occurrence period = 31.6 min |
| 2013 | Sun-Hee Kim et al. [57] | Bonn University Dataset | N/M | Detection of special characteristics | N/M | coercively adjusted auto regression (CA-AR) | Root mean square error = 0.029 |
| 2013 | Yang Zheng et al. [58] | European Epilepsy Database | Artefact removal and filtering | Bivariate empirical mode decomposition, Mean Phase Coherence (MPC) | A quantitative method based on the seizure prediction characteristic was proposed for the feature selection |
The preictal changes of the MPC time courses were used to raise the seizure alarms |
Sensitivity = 70-80% with Seizure Prediction Horizon = 10 min and False Prediction Rate(max) = 0.15 FP/h. |
| 2014 | Zhen Zhang et al. [59] | Real-time personal dataset | Optimal channel selection | Approximate entropy | N/M | N/M | Prediction accuracy = 94.59% False prediction rate = 0.084/h Mean prediction time = 26.64 min |
| 2014 | Nilufer Ozdemir et al. [60] | European Epilepsy Database | Artefact Removal and Segmentation | Hilbert Huang Transform, total energy | Filtering feature selection and Correlation-based Feature Selection (CFS) with the best first search algorithm |
Bayesian network | Sensitivity = 96.55% Mean detection latency = 33.21 False positives per hour (FPs/h) = 0.21 Time spent in warning (FP%) = 13.896 min |
| 2015 | Hamidreza Namazi et al. [61] | Real-time personal dataset | Filtering | Hurst exponent and fractal dimension | N/M | N/M | Seizure occurrence period = 25.76 seconds |
| 2015 | Kohtaroh Edakawa et al. [62] | Real-time personal dataset | N/M | Phase–amplitude coupling (PAC) |
θ -high γ, α -high γ, β -high γ, θ phase of 10–80 Hz amplitude, High γ amplitude alone |
Synchronisation index (SI) | Sensitivity = 100, False detection rate per hour = 0.713 |
| 2016 | A. Sharmila et al. [63] | Bonn University Dataset | N/M | Statistical features from Discrete Wavelet Transform (DWT) | N/M | Naïve Bayes, K-nearest neighbor | Accuracy = 100% (with Naïve Bayes) |
| 2016 | Mark H. Myers et al. [64] | CHB-MIT | Filtering | Phase Locking Thresholds | N/M | N/M | Sensitivity = 77% Precision = 88% False positive per hour = 0.17 |
| 2017 | Turky N. Alotaiby et al. [65] | CHB-MIT | N/M | Common Spatial Pattern (CSP) | N/M | Linear discriminant analysis (LDA) | Average sensitivity = 0.89 Average specificity = 0.37 Average False Prediction Rate = 0.39 Average prediction time = 68.71 minutes |
| 2017 | Amirmasoud Ahmadi et al. [66] | Bonn University Dataset | N/M | Wavelet packets transform | Standard Deviation, Root mean square | SVM with the radial basis function | Accuracy = 97.85% |
| 2018 | Ahmed I. Sharaf et al. [67] | Bonn University Dataset | Tunable Q-Wavelet Transformation (TQWT) | Chaotic features, statistical features, power spectrum features, Co-occurrence matrix |
Firefly algorithm | Random forest | Accuracy = 99% Precision = 97% Specificity = 97% Recall = 98% F-measure = 98% Matthew’s correlation coefficient = 95% |
| 2019 | Naghmeh Mahmoodian et al. [68] | European Epilepsy Database | Filtering and windowing | Cross-bispectral analysis | N/M | SVM | Sensitivity = 100% False positive rate (FPR) = 0.044 Prediction time = 51-96 minutes |
| 2019 | Agustina Garcés Correa et al. [69] | CHB-MIT | Adaptive filter and signal averaging | N/M | N/M | N/M | Sensitivity = 90.29% Specificity = 73.7% |
| 2019 | Hafeez A. Agboola et al. [70] | CHB-MIT | N/M | Low-level feature extraction | High level feature extraction | SVM, ANN | Sensitivity = 87.26% (SVM), 75.5% (ANN) False alarm per hour = 0.09 (SVM), 0.13 (ANN) Seizure occurrence period = 31 min (SVM), 29 min (ANN) |
| Year | Research Group | Dataset used | Preprocessing | Feature Extraction | Feature selection | Classifier used | Performance |
|---|---|---|---|---|---|---|---|
| 2005 | Nihal Fatma Guler et al. [71] | Bonn University database | N/M | Lyapunov exponents | N/M | RNN | Sensitivity = 96.88-96.13% Specificity = 97.38% Accuracy = 96.79% |
| 2015 | Khalid Abualsaud et al. [72] | Bonn University database | Compressive Sensing (CS), Discrete Cosine Transform (DCT) |
DWT | N/M | Noise-aware Signal Combination (NSC) ensemble classifier | Accuracy = 80% (for SNR=1dB), 84% (for SSNR=5dB), 88% (for SNR=10dB) |
| 2018 | Κostas М. Tsiouris et al. [73] | CHB-MIT | Segmentation | Cross-correlation, time domain, frequency domain, graph theory | N/M | LSTM | Sensitivity = 99.84% Specificity = 99.86% FPR per hour = 0.02 Preictal duration = 120 |
| 2018 | Punjal Agarwa et al. [74] | Kaggle | Image sampling, dimensionality reduction | FFT, CNN | N/M | Hybrid CNN-SVM | Accuracy = 97.07% Sensitivity = 96.47% Specificity = 98.81% |
| 2018 | Farrikh Alzami et al. [75] | Bonn University database | N/M | DWT | rank-aggregation (RA) | Adaptive hybrid feature selection-based ensemble |
Accuracy = 96-100% Sensitivity = 96.58-100% Specificity = 97.47-100% |
| 2018 | J.B. Schiratti et al. [76] | European Epilepsy Database | Down sampling | Time and frequency domain features | N/M | Logistic regression, Weighted Ensemble (WE) classifier |
ROC AUC score = 0.87 |
| 2018 | Lal Hussain [77] | Bonn University database | Wavelet threshold demonising method, Daudechies (db4) wavelet, PCA | Time domain, frequency domain and complexity | N/M | SVM, KNN, Decision tree, Ensemble | Accuracy = 99.5% (with SVM) AUC = 0.9991 (with SVM) |
| 2018 | Debdeep Sikdar et al. [78] | Bonn University database | Wavelet-based decomposition | Multifractal Detrended Fluctuation Analysis (MF-DFA) | N/M | SVM | Accuracy = 99.6% Precision = 99.3% Recall = 99.3% Specificity = 99.7 F-score = 99.3 |
| 2019 | Hisham Daoud et al. [22] | CHB-MIT | N/M | N/M | N/M | MLP DCNN+MLP DCNN+Bi-LSTM DCAE+Bi-LSTM DCAE+Bi-LSTM+CS |
Sensitivity = 99.72% Specificity = 99.60% Accuracy = 99.60% False Alarm per hour = 0.004 Prediction time = 1 hour |
| 2019 | Omer Turk et al. [79] | Bonn University database | N/M | Continuous Wavelet Transform (CWT) |
Resize image | CNN | Accuracy = 90.50-100% |
| 2020 | Yunyuan Gao et al. [80] | CHB-MIT | Signal denoising | Power spectrum density analysis | N/M | Inception-v3, ResNet152, Inception-ResNet-v2 | Accuracy = 92.6% Sensitivity = 97.1% Preictal duration (Minutes) = 30 |
7. DATASETS
Table 5 shows the comparative analysis of datasets used by various researchers for epileptic seizure detection and prediction. The datasets considered are highly unlike each other. The differences are in terms of EEG recording mechanism used i.e., sEEG or iEEG, number of subjects used, number of channels used, duration of the recordings, and number of recordings with the true positive case. The ‘*’ in the Table 5 suggests that the parameter has not been specified by the researchers. The research is also diversified in intra-subject (subject-specific), and inter-subject (adaptive) approaches. The selection of the dataset is highly dependent on these parameters. As derived from Table 5, majorly used datasets in the field of epileptic seizure detection/prediction are CHB-MIT [81, 82], Kaggle competition dataset [83], Bonn university dataset [84], and BCI competition dataset [85]. Apart from these, available significant datasets are the TUH dataset [86], European epilepsy dataset [87], and EEG epilepsy dataset [88]. Based on the subject-specific or adaptive approach of epileptic seizure prediction, the selection of the dataset is done [89]. For reproducibility of the results, utilization of the dataset must be clearly mentioned.
| References | Dataset | Type of EEG | No. of Subjects | No. of Channels |
|---|---|---|---|---|
| Syed Muhammad Usman et al. [4] | CHB-MIT | sEEG | 22 | 23 |
| Pouya Bashivan et al. [14] | Collected by authors | sEEG | 13 | 64 |
| Robin Tibor Schirrmeister et al. [15] | BCI competition-IV | sEEG | * | * |
| Sachin Talati et al. [16] | Bonn University | sEEG | * | 128/ single |
| Mohammad-Parsa Hosseini et al. [17] | BCI competition | iEEG | 9 | 15 |
| Mohammad-Parsa Hosseini et al. [18] | Kaggle competition | iEEG | 5 Dog and 2 Human | 16 and varying channels |
| Matthias Eberlein et al. [19] | Kaggle competition | iEEG | 5 Dog and 2 Human | 16 and varying channels |
| Isabell Kiral-Kornek et al. [20] | Cook et al. [89] | iEEG | 10 | 16 |
| David Ahmedt-Aristizabal et al. [21] | Bonn University | sEEG | * | 128/Single |
| Hisham Daoud et al. [22] | CHB-MIT | sEEG | 22 | 18/23 |
| Xinghua Yao et al. [23] | CHB-MIT | sEEG | 22 | 18/23 |
| Nick Hershey et al. [24] | Stanford hospital and Lucile Packard Children Hospital | sEEG & iEEG | 1,36,363 | 3-142 |
| Ghulam Muhammad et al. [25] | CHB-MIT | sEEG | 22 | 18/23 |
| Xiaoyan Wei et al. [26] | Xinjiang Medical University |
sEEG | 13 | * |
| Ali Emamia et al. [27] | NTT Medical Center Tokyo | sEEG | 16 | 19 |
| Turky N. Alotaiby et al. [65] | CHB-MIT | sEEG | 22 | 18/23 |
| P. Fergus et al. [90] | CHB-MIT | sEEG | 22 | 18/23 |
| Lung-Chang Lin et al. [40] | Kaohsiung Medical University Hospital |
sEEG | 5 | 21 |
| Julius Hulsmann [91] | PhysioNet | sEEG | 109 | 64 |
| Benjamin H. Brinkmann [92] | Kaggle competition | iEEG | 8 | 16 |
| Ricardo Aler [93] | BCI-III competition | sEEG | 3 | 32 |
| Ning Wang et al. [34] | Freiburg dataset | iEEG | 21 | 128 |
| Xinghua Yao et al. [51] | CHB-MIT | sEEG | 22 | 18/23 |
| Nipun Dilesh Perera et al. [94] | CHB-MIT | sEEG | 22 | 23 |
| Fayas Asharindavida et al. [95] | CHB-MIT, EPILEPSIAE-purchased | sEEG & iEEG | 22/125 | 22/217 |
| Yinxia Liu et al. [96] | Bonn University | sEEG | * | 128/Single |
| Gurwinder Singh et al. [97] | Bonn University | sEEG | * | 128/Single |
| I. Omerhodzic et al. [98] | Bonn University | sEEG | * | 128/Single |
| Punjal Agarwal et al. [74] | Kaggle competition | iEEG | 5 Dog and 2 Human | 16 and varying channels |
| Omer Turk et al. [79] | Bonn University | sEEG | * | 128/Single |
| Debdeep Sikdara et al. [78] | Bonn University | sEEG | * | 128/Single |
| Kostas M. Tsiouris et al. [73] | CHB-MIT | sEEG | 22 | 23 |
| Maarten Larmuseau et al. [99] | Kaggle competition | iEEG | 4 Dog | 16 |
| Kaat Vandecasteele et al. [100] | Hospital ECG, Wearable ECG, Wearable PPG | * | 11 | 1 |
| Ehsan Dadgar-kiani et al. [101] | Kaggle competition | iEEG | 1 | 16 |
| M. Stella Mercy [102] | Bonn University | iEEG | 2 | 1 |
| Sharanreddy et al. [103] | CHB-MIT | sEEG | 22 | 23 |
| U. Rajendra Acharya et al. [47] | Bonn University | sEEG | * | 128/Single |
| Ibrahim Aliyu et al. [52] | Bonn University | sEEG | * | 128/Single |
8. TOOLS AND LIBRARIES USED FOR IMPLEMENTATION
Though an ample amount of work has already been done in the field of epileptic seizure detection and prediction, the available published work does not contain much information related to the implementation environment used by the researchers. Table 6 summarizes the details available in past work on epileptic seizure detection and prediction.
It can be derived from Table 6 that most of the work is implemented using Matlab or Python. Various popular tools provided by Matlab are EEGLAB [104], Brainstorm [105], FieldTrip [106], EEGVIS [107], NFT [104], and BCILAB [104]. Python provides different libraries like MNE [108], MNE-Python [109], PyEEG [110], and Pyprep [111]. Whereas R provides packages like eegkit [112], eegUtils [113], eegR [114], erpR [115], and ERP [116]. While undertaking any research project which includes EEG signal analysis, researchers spent an ample amount of time in finalizing the platform to adapt for implementation. Following detail shows a comparison of Matlab, Python, and R concerning various parameters useful for implementation:
| References | Framework Used |
|---|---|
| Syed Muhammad Usman et al. [4] | Matlab |
| Pouya Bashivan et al. [14] | Lasagne |
| Mohammad-Parsa Hosseini et al. [18] | Pytorch |
| Xiaoyan Wei et al. [26] | Python with Tensorflow |
| Ali Emami et al. [27] | Python |
| Lung-Chang Lin et al. [40] | Weka |
| Julius Hu¨lsmann et al. [91] | Numpy, Scipy, MNE |
| Benjamin H. Brinkmann et al. [92] | LibSVM |
| Ricardo Aler et al. [93] | Weka |
| Ning Wang et al. [34] | Matlab spiderbox toolbox |
| Fayas Asharindavida et al. [95] | Matlab |
| Maarten Larmuseau [99] | Sci-kit learn, keras |
| Ehsan Dadgar-Kiani et al. [101] | Sci-kit learn, keras |
| M. Stella Mercy [102] | LibSVM |
| Sharanreddy et al. [103] | Matlab |
| U. Rajendra Acharya et al. [47] | Matlab |
| Ibrahim Aliyu et al. [52] | Python with Tensorflow and Keras |
Open Source: The major advantage of using Python and R over Matlab is that they are open source, making them quite attractive solution to many applications. Applications developed in Python and R can be widely distributable, making it easier to enable collaboration between scientists at various locations. Maturity of Tools: The tools supported by Matlab are quite mature to use compared to Python and R. It is best suitable for detailed analysis of the EEG signals. Flexibility of Manipulation: The maturity makes it less flexible to manipulate and extend features in Matlab. Community Support: Community support has important significance in selecting the specific platform for research. Matlab and Python both have huge community support when it comes to EEG signal processing. This area is quite evolving in R; however, the statistical analysis performed by R is no comparable.
One of the best solutions could be to use the best feature of each platform and use it in integration. Python has an Oct2py library [117], which converts Python data structures to Matlab or Octave data structures and vice versa. It is the simplest and most stable way to run Matlab functions on Python, and most EEGLAB functions may be called from within python using this method. R provides the reticulate package [118], which allows running Python code directly within R.
CONCLUSION
EEG-based epilepsy detection and prediction using a machine learning approached has taken a boost following the technology evolution. The essential aspects of EEG waveform-based research for epileptic seizure detection and prediction have been discussed in detail. The comparative analysis of various datasets and implementation platforms considered in the past approaches along with details of traditional machine learning, deep learning, and combination of both - the hybrid approach is given, which would help novice to begin. The selection of dataset and implementation environment is highly dependent on requirement of the research. Various criteria play an important role, such as for datasets: subject-specific research or generalized research to be carried out, number of channels to consider, number of patients to consider; and for implementation environment: budget, programming expertize, and size of the dataset. Shortcomings of epileptic seizure prediction approaches are also discussed to shed light on future enhancements.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none.


